Fine-Tuning FLAN-T5 for Biomedical Lay Summarisation (BioLaySumm 2025)
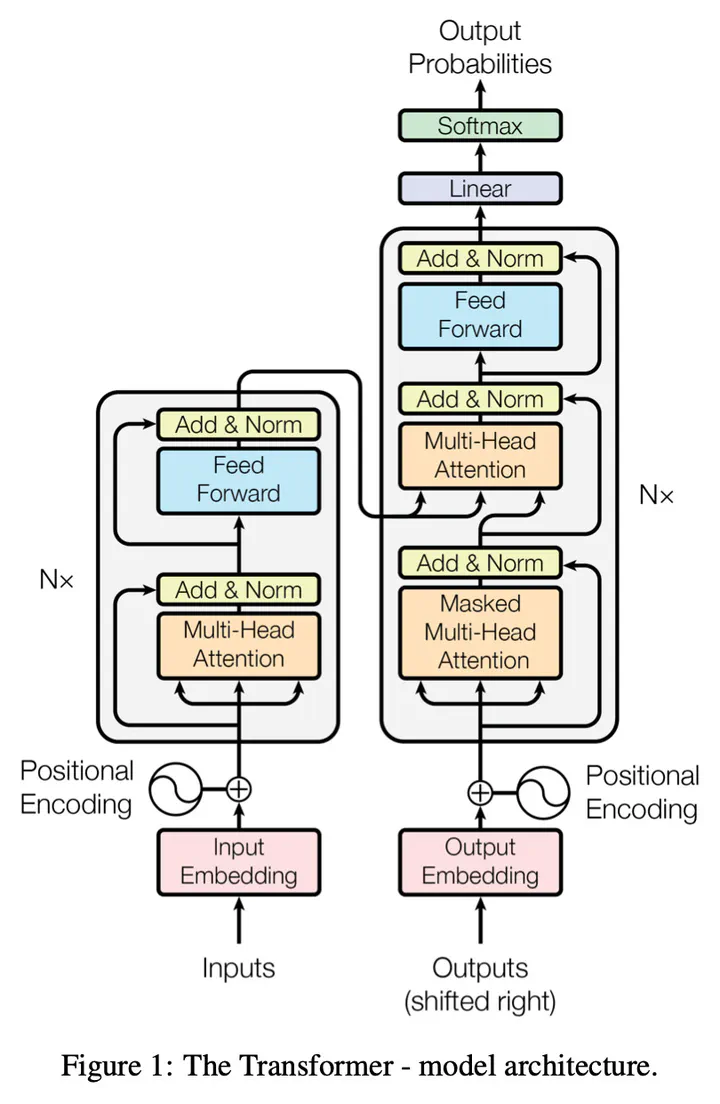 FLAN-T5 architecture, LoRA adapters, and training results.
FLAN-T5 architecture, LoRA adapters, and training results.For full methodology, experiments, training scripts, and results, see the GitHub repository.
Overview
This project focuses on biomedical lay summarisation, translating expert-level radiology reports into language accessible to non-experts. The work fine-tunes instruction-tuned FLAN-T5 models on the BioLaySumm 2025 dataset and systematically evaluates different adaptation strategies.
Methods
Three optimisation strategies are explored:
- Full Fine-Tuning (FFT): updates all model parameters.
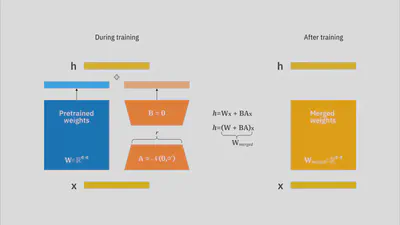
- LoRA (PEFT): updates a small set of low-rank adapter weights (~2–3% of parameters).
- Evolution Strategies (ES): gradient-free optimisation via population-based parameter perturbations.
Performance is evaluated using ROUGE-1/2/L/Lsum, with analysis of compute cost, convergence behaviour, and parameter efficiency.
Key Findings
- LoRA achieves comparable or slightly higher ROUGE scores than full fine-tuning while training only ~2–3% of parameters.
- FLAN-T5-Base + LoRA provides the best balance of quality and efficiency.
- Evolution Strategies offer fast iteration but underperform gradient-based methods for this task.
Example Visuals
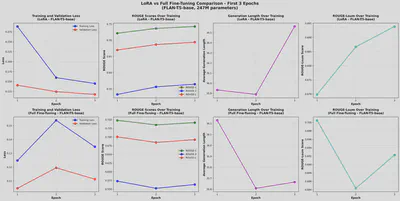

Reproducibility & Code
All experiments were run on NVIDIA A100 GPUs using PyTorch and Hugging Face Transformers. Training scripts, hyperparameters, datasets, and seeds are fully documented in the repository.
👉 Full code and documentation:
https://github.com/Jevi-Waugh/BioLaySumm-Flan-T5/tree/topic-recognition/recognition/FLAN-T5-Jevi-Waugh